|
Cytoskeleton Inc
ubiquitinated protein Ubiquitinated Protein, supplied by Cytoskeleton Inc, used in various techniques. Bioz Stars score: 95/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/ubiquitinated protein/product/Cytoskeleton Inc Average 95 stars, based on 1 article reviews
ubiquitinated protein - by Bioz Stars,
2026-02
95/100 stars
|
Buy from Supplier |
|
MedChemExpress
recombinant chip protein Recombinant Chip Protein, supplied by MedChemExpress, used in various techniques. Bioz Stars score: 92/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/recombinant chip protein/product/MedChemExpress Average 92 stars, based on 1 article reviews
recombinant chip protein - by Bioz Stars,
2026-02
92/100 stars
|
Buy from Supplier |
|
R&D Systems
recombinant ub Recombinant Ub, supplied by R&D Systems, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/recombinant ub/product/R&D Systems Average 96 stars, based on 1 article reviews
recombinant ub - by Bioz Stars,
2026-02
96/100 stars
|
Buy from Supplier |
|
Sino Biological
ubiquitin Ubiquitin, supplied by Sino Biological, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/ubiquitin/product/Sino Biological Average 94 stars, based on 1 article reviews
ubiquitin - by Bioz Stars,
2026-02
94/100 stars
|
Buy from Supplier |
|
R&D Systems
tetra ubiquitin  Tetra Ubiquitin, supplied by R&D Systems, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/tetra ubiquitin/product/R&D Systems Average 93 stars, based on 1 article reviews
tetra ubiquitin - by Bioz Stars,
2026-02
93/100 stars
|
Buy from Supplier |
|
R&D Systems
ub-rh110  Ub Rh110, supplied by R&D Systems, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/ub-rh110/product/R&D Systems Average 93 stars, based on 1 article reviews
ub-rh110 - by Bioz Stars,
2026-02
93/100 stars
|
Buy from Supplier |
|
Boster Bio
microfluidic chip  Microfluidic Chip, supplied by Boster Bio, used in various techniques. Bioz Stars score: 92/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/microfluidic chip/product/Boster Bio Average 92 stars, based on 1 article reviews
microfluidic chip - by Bioz Stars,
2026-02
92/100 stars
|
Buy from Supplier |
|
Proteintech
nedd8 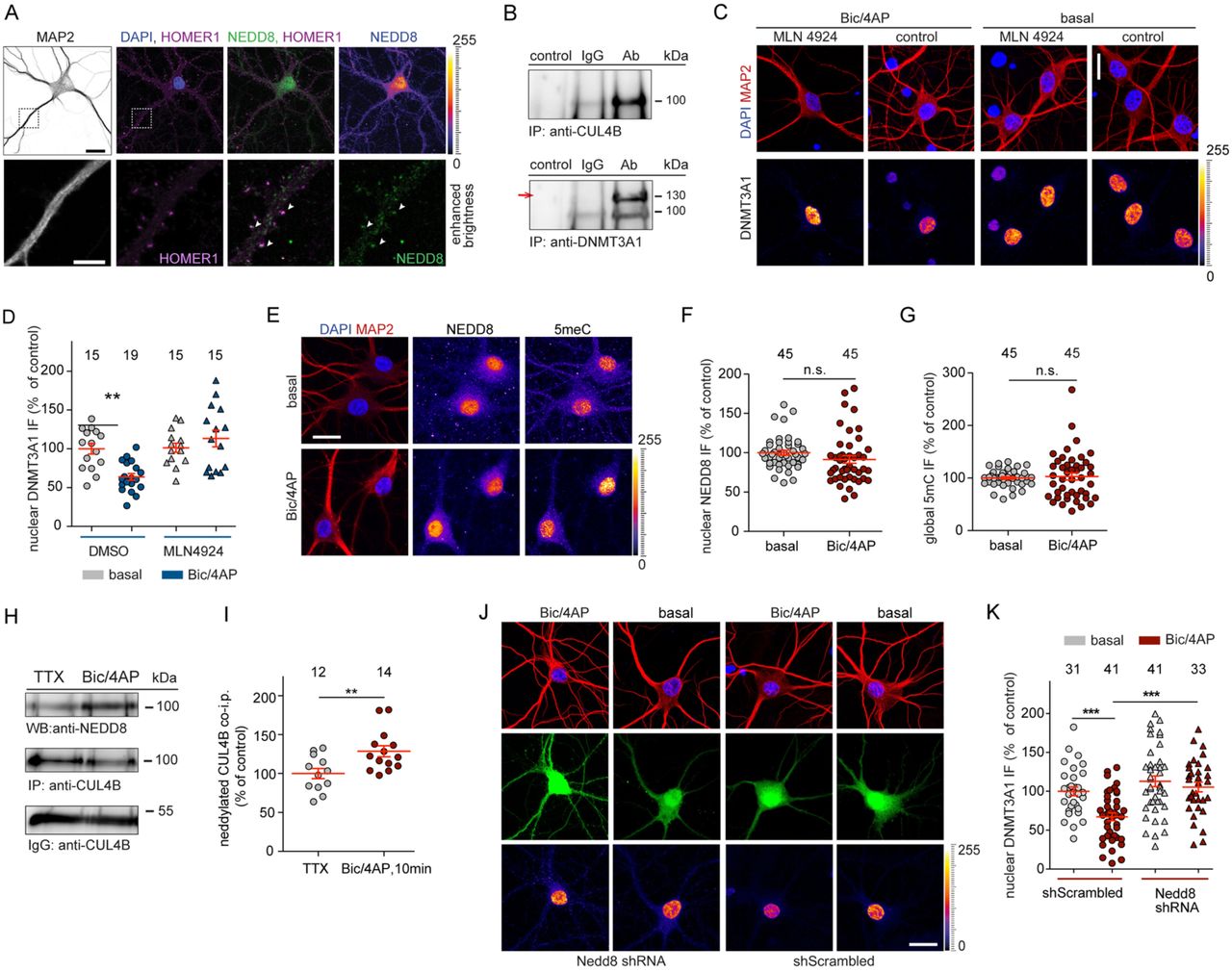 Nedd8, supplied by Proteintech, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/nedd8/product/Proteintech Average 93 stars, based on 1 article reviews
nedd8 - by Bioz Stars,
2026-02
93/100 stars
|
Buy from Supplier |
|
Proteintech
atg12  Atg12, supplied by Proteintech, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/atg12/product/Proteintech Average 93 stars, based on 1 article reviews
atg12 - by Bioz Stars,
2026-02
93/100 stars
|
Buy from Supplier |
|
Proteintech
mouse anti sumo2 3  Mouse Anti Sumo2 3, supplied by Proteintech, used in various techniques. Bioz Stars score: 95/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/mouse anti sumo2 3/product/Proteintech Average 95 stars, based on 1 article reviews
mouse anti sumo2 3 - by Bioz Stars,
2026-02
95/100 stars
|
Buy from Supplier |
|
Proteintech
anti sumo1 antibody  Anti Sumo1 Antibody, supplied by Proteintech, used in various techniques. Bioz Stars score: 95/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/anti sumo1 antibody/product/Proteintech Average 95 stars, based on 1 article reviews
anti sumo1 antibody - by Bioz Stars,
2026-02
95/100 stars
|
Buy from Supplier |
|
Proteintech
ubl4a  Ubl4a, supplied by Proteintech, used in various techniques. Bioz Stars score: 92/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/ubl4a/product/Proteintech Average 92 stars, based on 1 article reviews
ubl4a - by Bioz Stars,
2026-02
92/100 stars
|
Buy from Supplier |
Image Search Results

Journal: Biochemical pharmacology
Article Title: Platinum-containing compound platinum pyrithione is stronger and safer than cisplatin in cancer therapy
doi: 10.1016/j.bcp.2016.06.019
Figure Lengend Snippet: PtPT inhibits proteasome function in multiple cancer cells. (A and B) Dose-dependent accumulation of proteasomal substrates in K562, A549 and A549/DDP cells. These cells were treated with PtPT (2.5, 5, 7.5, 10 μM), bortezomib/Velcade (Vel, 50 nM), CDDP (20 μM), or b-AP15 (0.5 μM) for 12 h, followed by detecting the levels of total (Ub-prs) and K48-linked (K48-) ubiquitin conjugates, p27, and PARP cleavage with western blot analysis. (C and D) The changes of a surrogate proteasome substrate (GFPu). HEK-293 cells stably expressing GFPu were treated with PtPT for 12 h, then ubiquitinated proteins and GFPu protein were detected with western blot or imaged under an inverted fluorescence microscope (scale bar, 50 μm). Bortezomib/Velcade (Vel) and b-AP15 were used as positive controls. CDDP was used as a negative control. One representative blot (A/B/D, upper panel) out of three independent experiments is shown and quantified by densitometric analysis (lower panel). Values are expressed as mean ± SD ( n = 3). * P < 0.05, compared with each control.
Article Snippet: PtPT was synthesized in our lab. Other agents are bortezomib (BD Biosciences, San Jose, CA); NEM, Penicillin, Streptomycin, Cisplatin (Sigma–Aldrich Inc., St. Louis, MO); b-AP15, Suc-Leu-Leu-V al-Tyr-aminomethylcoumarin (Suc-LLVY-AMC), Z-Leu-Leu-Glu-AMC (Z-LLE-AMC), Boc-Leu-Arg-Arg-AMC (Boc-LRR-AMC), 19S, 20S and 26S human proteasomes, HA-Ubiquitin-Vinyl Sulfone (HA-Ub-VS), K48-linked
Techniques: Western Blot, Stable Transfection, Expressing, Fluorescence, Microscopy, Negative Control

Journal: Biochemical pharmacology
Article Title: Platinum-containing compound platinum pyrithione is stronger and safer than cisplatin in cancer therapy
doi: 10.1016/j.bcp.2016.06.019
Figure Lengend Snippet: PtPT inhibits proteasomal deubiquitinase (DUB) USP14 and UCHL5. (A) The effect of PtPT on cytoplasmic total DUB activities. A549 cell lysates were exposed to PtPT (5.0 μM) and dynamic DUB activity was measured. NEM was used as a positive control. (B) The effect of PtPT at the indicated dosages on the activity of 5 recombinant non-proteasomal deubiquitinases (UCHL1, UCHL2, BAP1, USP5, and USP7) was tested. End-point DUB activity is shown. n = 3, Mean ± SD. (C) The effect of PtPT on proteasomal DUB activities. Purified 26S proteasomes were incubated with PtPT or CDDP (5.0 μM), and then DUB activity kinetics were measured. NEM was used as a positive control. (D) Purified 26S proteasomes were treated with PtPT (0.5, 1.0, 2.0 μM), then dynamic DUB activity was recorded. (E) Ubiquitin chain disassembly assay. The disassembly of K48-linked ubiquitin tetramers mediated by the 26S proteasome was detected in the presence of PtPT or b-AP15.One representative blot (E/left panel) out of three independent experiments is shown and quantified by densitometric analysis (E/right panel). Values are expressed as mean ± SD ( n = 3). * P < 0.05, compared with each Ub4. (F) Computational molecular docking of PtPT with UCHL5 and USP14 of the 26S proteasomes. The following data are shown: the structure of platinum pyrithione (L1); the structure of platinum pyrithione intermediate (L2); the binding modes of compound L2 at the active site of USP14; the binding models of compound L2 at the active site of UCHL5. (G) Active-site-directed labeling of proteasomal DUBs. Purified 26S proteasomes were treated with PtPT, NEM, or b-AP15, followed by labeling with HA-UbVS and immunoblotting for HA. One representative blot (E/left panel and G/upper panel) out of three independent experiments is shown and quantified by densitometric analysis (E/right panel and G/lower panel). Values are expressed as mean ± SD ( n = 3). * P < 0.05, compared with each 26S.
Article Snippet: PtPT was synthesized in our lab. Other agents are bortezomib (BD Biosciences, San Jose, CA); NEM, Penicillin, Streptomycin, Cisplatin (Sigma–Aldrich Inc., St. Louis, MO); b-AP15, Suc-Leu-Leu-V al-Tyr-aminomethylcoumarin (Suc-LLVY-AMC), Z-Leu-Leu-Glu-AMC (Z-LLE-AMC), Boc-Leu-Arg-Arg-AMC (Boc-LRR-AMC), 19S, 20S and 26S human proteasomes, HA-Ubiquitin-Vinyl Sulfone (HA-Ub-VS), K48-linked
Techniques: Activity Assay, Positive Control, Recombinant, Purification, Incubation, Binding Assay, Labeling, Western Blot

Journal: Biochemical pharmacology
Article Title: Platinum-containing compound platinum pyrithione is stronger and safer than cisplatin in cancer therapy
doi: 10.1016/j.bcp.2016.06.019
Figure Lengend Snippet: Proteasome inhibition is required for PtPT to induce apoptosis. (A and B) PtPT time-dependently induces proteasome function inhibition and apoptosis in cancer cells (A549, A549/DDP and K562). The cells in culture treated with PtPT (5 μM) were collected at the indicated time points for western blot analyses for ubiquitinated proteins including total ubiquitin conjugates (Ub-prs) and K48-linked polyubiquitins, p27, as well as cleaved PARP. (C) An illustration of the coordination of Pt with ethylenediaminetetraacetic acid (EDTA) to inactivate PtPT. (D and E) EDTA prevented most PtPT-induced proteasome inhibition and apoptosis. A549 cells were treated with PtPT with/without chelating agent EDTA (100 μM) for 12 h, total ubiquitin conjugates and PARP were detected with western blot analysis, and apoptosis cells were recorded with PI staining followed by imaging under fluorescence microscopy. One representative blot (A/B/D, upper panel) out of three independent experiments is shown and quantified by densitometric analysis (lower panel). Values are expressed as mean ± SD ( n = 3). * P < 0.05, compared with each control.
Article Snippet: PtPT was synthesized in our lab. Other agents are bortezomib (BD Biosciences, San Jose, CA); NEM, Penicillin, Streptomycin, Cisplatin (Sigma–Aldrich Inc., St. Louis, MO); b-AP15, Suc-Leu-Leu-V al-Tyr-aminomethylcoumarin (Suc-LLVY-AMC), Z-Leu-Leu-Glu-AMC (Z-LLE-AMC), Boc-Leu-Arg-Arg-AMC (Boc-LRR-AMC), 19S, 20S and 26S human proteasomes, HA-Ubiquitin-Vinyl Sulfone (HA-Ub-VS), K48-linked
Techniques: Inhibition, Western Blot, Staining, Imaging, Fluorescence, Microscopy

Journal: Biochemical pharmacology
Article Title: Platinum-containing compound platinum pyrithione is stronger and safer than cisplatin in cancer therapy
doi: 10.1016/j.bcp.2016.06.019
Figure Lengend Snippet: PtPT inhibits tumor growth and proteasome function in vivo . (A–D) BALB/c nude mice bearing A549 tumors were treated with vehicle or PtPT (5 mg/kg/day, i.p.) or CDDP (2.5 mg/kg/2 day, i.p.) for 15 days. Tumor size was recorded every other day. Tumor size (A), tumor images (B), tumor weight (C) and body weight (D) are shown. n = 6, mean ± SD. * P < 0.05, ** P < 0.01, compared with each control. (E) Liver function was assessed by detecting serum AST and ALT. n = 6, mean ± SD. * P < 0.05, compared with each control. AU, arbitrary unit. (F–H) BALB/c nude mice were treated with a single injection of vehicle, PtPT (15 mg/kg, i.p.) or CDDP (15 mg/kg/, i.p.), and sacrificed 4 days later. The graph showing the serum BUN (F) and creatinine (G) levels in each group. * P < 0.05, compared to vehicle control ( n = 6). Representative photomicrographs (200×) of mouse kidney sections with Periodic Acid Schiff staining are shown (H). (I) Representative immunohistochemical staining results are shown for total (Ub-prs) or K48-linked ubiquitin accumulation, proteasome substrate protein (p21), and cleaved caspase-3 in A549 tumor tissues from nude mice treated with vehicle, PtPT, or CDDP (200×). (J–M) BALB/c nude mice bearing K562 tumors were treated with either vehicle or PtPT (7.5 mg/kg/day, i.p.) for consecutive 15 days, Tumor size was recorded every other day. Tumor size (G), tumor images (H), tumor weight (I) and body weight (J) are shown. n = 6, mean ± SD. ** P < 0.01, compared with each control. (N) Representative images of immunohistochemical staining for total (Ub-prs), K48-linked ubiquitin accumulation, proteasome substrate protein (p27), and cleaved caspase-3 in tumor tissues from mice bearing K562 tumors treated with either vehicle or PtPT (200×).
Article Snippet: PtPT was synthesized in our lab. Other agents are bortezomib (BD Biosciences, San Jose, CA); NEM, Penicillin, Streptomycin, Cisplatin (Sigma–Aldrich Inc., St. Louis, MO); b-AP15, Suc-Leu-Leu-V al-Tyr-aminomethylcoumarin (Suc-LLVY-AMC), Z-Leu-Leu-Glu-AMC (Z-LLE-AMC), Boc-Leu-Arg-Arg-AMC (Boc-LRR-AMC), 19S, 20S and 26S human proteasomes, HA-Ubiquitin-Vinyl Sulfone (HA-Ub-VS), K48-linked
Techniques: In Vivo, Injection, Staining, Immunohistochemical staining

Journal: bioRxiv
Article Title: Synaptic control of DNA-methylation involves activity-dependent degradation of DNMT3a1 in the nucleus
doi: 10.1101/602151
Figure Lengend Snippet: A NEDD8 is abundantly expressed in the nucleus of 16 DIV hippocampal neurons whereas weaker staining was observed at synapses. Scale bar 20 µm (top panel), 5 µm (lower panel). B Immunoprecipitation of CUL4B protein from nuclear extracts of cortical primary neurons (top panel) results in co-precipitation of Dnmt3a1 (lower panel, indicated with an arrow). C and D Hippocampal primary neurons were treated with bic/4AP for 6 h in the presence or absence of MLN4924 (5 nM). Scale bars, 20 µm. Quantitative immunocytochemistry revealed that blocking neddylation prevents Dnmt3a1 degradation. E Representative immunofluorescence images of nuclear NEDD8 and total cytosine methylation (5mC) at basal conditions or after treatment with bic/4AP for 6 h. Scale bar is 20 µm. F and G Upon synaptic activation nuclear NEDD8 and 5mC levels remained unchanged. Student’s t-test. n.s. not significant. H and I CUL4B was immunoprecipitated from nuclear extracts of primary cortical neurons and neddylated CUL4B was quantified. Following 10 min of synaptic stimulation the amount of neddylated CUL4B was increased. Student’s t-test **p<0.01. J and K shRNA-based knockdown of NEDD8 in hippocampal primary neurons occluded the DNMT3A1 degradation following 10 min-long bic/4AP treatment and fixation of cells 3 h after washout of the drug-containing-media. Two-way ANOVA followed by Bonferroni’s post-hoc test. ***p<0.001, scale bar, 20 µm.
Article Snippet: The following antibodies were used in this study: DNMT3AN (1:2000) and DNMT3A-mid (1:2000), CUL4B (1:1000, Proteintech, catalog #12916-1-AP),
Techniques: Staining, Immunoprecipitation, Immunocytochemistry, Blocking Assay, Immunofluorescence, Methylation, Activation Assay, shRNA, Knockdown

Journal: bioRxiv
Article Title: Synaptic control of DNA-methylation involves activity-dependent degradation of DNMT3a1 in the nucleus
doi: 10.1101/602151
Figure Lengend Snippet: A Heterologous expression of GFP-DNMT3A1 and empty myc-vector or human myc-Cullins (CUL1, CUL2, CUL3, CUL4A, CUL4B, CUL5, CUL7) in HEK293-T cells. DNMT3A1 was immunoprecipitated from total cell extracts using anti-GFP antibodies coupled to microbeads. Cullin proteins were detected in the immunoprecipitate with a myc-antibody. Same volume of IP and input per sample were loaded. B HEK-T cells were transfected with expression vectors for GFP-DNMT3A1 and myc-CUL4B or myc-vector. DNMT3A1 was immunoprecipitated from total cell extracts using anti-GFP antibodies coupled to microbeads. CUL4B was detected in the immunoprecipitate with a myc-antibody. C and D Poly-ubiquitination of DNMT3A1 in the immunoprecipitate is enhanced when DNMT3A1 was co-expressed with CUL4B. Unpaired two-tailed Student’s t-test, *p<0.05. E shRNA knockdown of CUL4B reduces polyubiquitination of DNMT3A1. F and G HEK-T cells were transfected with expression vectors for GFP-DNMT3A1, myc-CUL4B ( F ) and also with active HA-Nedd8 ( G ). Total cell extracts were immunoprecitated using anti-GFP antibodies. ( F ) Less DNMT3A1 ubiquitination was observed when the NAE inhibitor MLN4924 (94nM) was applied for 24h. ( G ) Neddylated CUL4B was detected with anti-HA antibody in the DNMT3A1 elute.
Article Snippet: The following antibodies were used in this study: DNMT3AN (1:2000) and DNMT3A-mid (1:2000), CUL4B (1:1000, Proteintech, catalog #12916-1-AP),
Techniques: Expressing, Plasmid Preparation, Immunoprecipitation, Transfection, Ubiquitin Proteomics, Two Tailed Test, shRNA, Knockdown

Journal: bioRxiv
Article Title: Synaptic control of DNA-methylation involves activity-dependent degradation of DNMT3a1 in the nucleus
doi: 10.1101/602151
Figure Lengend Snippet: A and B DIV 14 hippocampal neurons were transfected with RFP expressed under control of an actin promoter and cells were fixed two days after transfection. Homer1 positive dendritic spine numbers were counted. There is not any significant alteration in the total spine number following MLN4924 treatments following the synaptic stimulation with bic/4AP for 6 h. Scale bars are 5 µm. C and D Representative images and quantification display the endogenous rat-Nedd8 knockdown generated in 16 DIV primary hippocampal neurons. Unpaired two-tailed Student’s t-test, *** p<0.0001. Scale bar in C is 20 µm. E Heterologous expression of human-NEDD8 and shRNA mRNA knockdown and shScrambled control in HEK-T cells was detected with either the anti-myc antibody (upper panel) or the anti-NEDD8 antibody (lower panel). The knockdown of Nedd8 was detected by anti-myc and as well as anti-NEDD8 antibodies.
Article Snippet: The following antibodies were used in this study: DNMT3AN (1:2000) and DNMT3A-mid (1:2000), CUL4B (1:1000, Proteintech, catalog #12916-1-AP),
Techniques: Transfection, Control, Knockdown, Generated, Two Tailed Test, Expressing, shRNA

Journal: iScience
Article Title: MBNL2 promotes aging-related cardiac fibrosis via inhibited SUMOylation of Krüppel-like factor4
doi: 10.1016/j.isci.2024.110163
Figure Lengend Snippet: Inhibition of MBNL2 activated SUMOylation of KLF4 (A–D) Representative graphs of SUMO1/2/3 protein in Young vs. Aged, Control vs. H 2 O 2 , oe-NC vs. oe-MBNL2, sh-NC vs. sh-MBNL2. (E–G) Representative and statistical graphs of SENP1 and SUMO1 protein. (H–J) Representative and statistical graphs of KLF4 and SENP1 protein. (K) Representative immunoblots of KLF4-SUMOylation after incubation with sh-MBNL2 or/and H 2 O 2 in CFs ( n = 3). (L–O) Western blot analysis and quantification of TGF-β1, T-SMAD3 and P-SMAD3 protein levels (n = 3–6, data are expressed as mean ± SEM, ∗ p < 0.05, ∗∗ p < 0.01 vs. the control group, # p < 0.05, ## p < 0.01 vs. the H 2 O 2 group, & p < 0.05, && p < 0.01 vs. the H 2 O 2 +sh-MBNL2 group).
Article Snippet:
Techniques: Inhibition, Control, Western Blot, Incubation

Journal: iScience
Article Title: MBNL2 promotes aging-related cardiac fibrosis via inhibited SUMOylation of Krüppel-like factor4
doi: 10.1016/j.isci.2024.110163
Figure Lengend Snippet:
Article Snippet:
Techniques: Recombinant, Staining, cDNA Synthesis, SYBR Green Assay, Magnetic Beads, Plasmid Preparation, Extraction, Microarray, Software

Journal: Experimental Animals
Article Title: Ubiquitin-like 4A alleviates the progression of intracerebral hemorrhage by regulating oxidative stress and mitochondrial damage
doi: 10.1538/expanim.24-0035
Figure Lengend Snippet: (a) Schematic diagram showing the experimental procedure of the ICH rat. (b) Forelimb placing test and corner turn test of rats. (c) Brain water content of rats. (d) Real-time PCR and western blot were used to verify the expression of UBL4A in brain tissue. (e) Typical hematoxylin-eosin (HE) staining images presenting the striatal hematomas in rats. Scale bar: 500 µ m (left) and 100 µ m (right). (f) Fluoro-Jade B (FJB) staining was performed to detect neuronal cell death in rats. Scale bar: 50 µ m. (g) Co-localization of ubiquitin-like protein 4A (UBL4A) and NeuN was detected by immunofluorescence staining. Scale bar: 50 µ m. ICH, intracerebral hemorrhage. * P <0.05, ** P <0.01, ns: no significance.
Article Snippet: Tissue sections were blocked with
Techniques: Real-time Polymerase Chain Reaction, Western Blot, Expressing, Staining, Ubiquitin Proteomics, Immunofluorescence

Journal: Experimental Animals
Article Title: Ubiquitin-like 4A alleviates the progression of intracerebral hemorrhage by regulating oxidative stress and mitochondrial damage
doi: 10.1538/expanim.24-0035
Figure Lengend Snippet: (a) The diagram depicts the flow chart of the cell experiment. (b) Real-time PCR and western blot were used to verify the expression level of ubiquitin-like protein 4A (UBL4A) in neurons. (c) CCK-8 assay was performed to determine cell viability. (d) TUNEL staining was used to detect cell apoptosis. Scale bar: 50 µ m. LV-shNC, negative control shRNA; LV-shUBL4A, shUBL4A knockdown; LV-vector, empty vector; LV-UBL4A, UBL4A overexpression. * P <0.05, ** P <0.01.
Article Snippet: Tissue sections were blocked with
Techniques: Real-time Polymerase Chain Reaction, Western Blot, Expressing, Ubiquitin Proteomics, CCK-8 Assay, TUNEL Assay, Staining, Negative Control, shRNA, Knockdown, Plasmid Preparation, Over Expression

Journal: Experimental Animals
Article Title: Ubiquitin-like 4A alleviates the progression of intracerebral hemorrhage by regulating oxidative stress and mitochondrial damage
doi: 10.1538/expanim.24-0035
Figure Lengend Snippet: (a) The level of reactive oxygen species (ROS) in cells was determined by flow cytometry using a dichlorodihydrofluorescein diacetate (DCFH-DA) based kit (cell number: ×10 3 ). (b) Manganese superoxide dismutase (MnSOD) activity was measured with MnSOD assay kit. (c) Detection of mitochondrial membrane potential by JC-1 staining. (d) The relative mitochondrial DNA (mtDNA) level was detected by real-time PCR. LV-shNC, negative control shRNA; LV-shUBL4A, shUBL4A knockdown; LV-vector, empty vector; LV-UBL4A, UBL4A overexpression. * P <0.05, ** P <0.01.
Article Snippet: Tissue sections were blocked with
Techniques: Flow Cytometry, Activity Assay, Membrane, Staining, Real-time Polymerase Chain Reaction, Negative Control, shRNA, Knockdown, Plasmid Preparation, Over Expression

Journal: Experimental Animals
Article Title: Ubiquitin-like 4A alleviates the progression of intracerebral hemorrhage by regulating oxidative stress and mitochondrial damage
doi: 10.1538/expanim.24-0035
Figure Lengend Snippet: (a) Real-time PCR to verify the expression of miR-34a-5p in neurons. (b) Real-time PCR and western blot were used to verify the expression level of ubiquitin-like protein 4A (UBL4A) in neurons with miR-34a-5p knockdown. (c) TUNEL staining was used to detect cell apoptosis. Scale bar: 50 µ m. (d) The level of reactive oxygen species (ROS) in cells was determined by flow cytometry using a dichlorodihydrofluorescein diacetate (DCFH-DA) based kit (cell number: ×10 3 ). LV-shNC, negative control shRNA; LV-miR-34a-sponge, miR-34a-5p knockdown. ** P <0.01.
Article Snippet: Tissue sections were blocked with
Techniques: Real-time Polymerase Chain Reaction, Expressing, Western Blot, Ubiquitin Proteomics, Knockdown, TUNEL Assay, Staining, Flow Cytometry, Negative Control, shRNA

Journal: Experimental Animals
Article Title: Ubiquitin-like 4A alleviates the progression of intracerebral hemorrhage by regulating oxidative stress and mitochondrial damage
doi: 10.1538/expanim.24-0035
Figure Lengend Snippet: (a) Predicted miR-34a-5p target sequences in 3′-UTR of Ubl4a (Left). Dual-luciferase reporter assay was conducted to verify whether ubiquitin-like protein 4A (UBL4A) was the target of miR-34a-5p (Right). NC mimics, negative control mimics; WT, wild type; MUT, mutant type. (b) Western blot was used to verify the expression level of UBL4A in neurons. (c) TUNEL staining was used to detect cell apoptosis. Scale bar: 50 µ m. (d) The level of reactive oxygen species (ROS) in cells was determined by flow cytometry using a dichlorodihydrofluorescein diacetate (DCFH-DA) based kit (cell number: ×10 3 ). LV-shNC, negative control shRNA; LV-miR-34a-sponge, miR-34a-5p knockdown. LV-shUBL4A, shUBL4A knockdown. * P <0.05, ** P <0.01.
Article Snippet: Tissue sections were blocked with
Techniques: Luciferase, Reporter Assay, Ubiquitin Proteomics, Negative Control, Mutagenesis, Western Blot, Expressing, TUNEL Assay, Staining, Flow Cytometry, shRNA, Knockdown